INTRODUCTION
The explosion of information is posing a challenge to the existing means for data storage. It is anticipated that current storage methods (ex. magnetic and optical media) will be inadequate to store the exponentially growing data. In 2020, every individual in the world had generated around 1.7 MB of data each second, which amounted to 418 zettabytes, requiring approximately 418 billion one-terabyte hard drives for storage.[1] To provide a perspective, the Large Hadron Collider, located at the European Organization for Nuclear Research (CERN), is an example of a contributor to the unprecedented research data that is being added continuously. It generates about 50 million GB of data per year as it records the results of experiments involving approximately 600 million particle collisions per second. In the area of life sciences, DNA sequencing alone generates millions of GB of data per year; and it is predicted that within a decade, we will be swamped with 40 billion GB of genomic data.
Traditional mass-storage technologies are starting to approach their exhaustive limits while the need for data storage keeps surging. With hard-disk drives, there is a limit of 1 TB per square inch.[2] The biggest tape archive facilities can store an exabyte of data, but these facilities take up ample space, cost in billions for upkeep, use considerable amounts of energy, and are required to be copied at regular intervals to ensure that the data is not lost due to degradation.[3] Also temperature fluctuations can induce the magnetically charged material of the disk to flip, thus corrupting the data it holds. Better heat-resistant material needs to be utilized for which technology has to be altered. This, in turn, would require huge investments.[2] Therefore, data scientists are looking for better, more stable, and space-efficient alternatives to store huge datasets. DNA-based data storage has recently emerged as a promising approach for long-term digital information storage. Highly condensed DNA has great potential to become a storage material of the future.
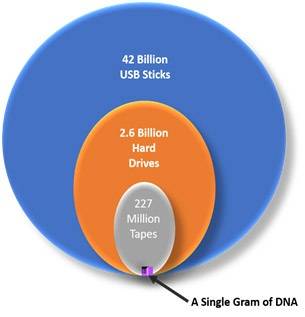
A solution to this issue of requirement of digital storage space may be found in deoxyribonucleic acid (DNA), the molecular repository of biological information. DNA has an astonishing ability to store biological data, as it is the basic unit of storage system for all the information that governs biological life. DNA is not only abundant and sustainable, it also provides greater storage density than the currently available data storage media.[4] Comparison of amount of traditional data storage systems versus DNA required to store 40 ZB data has been provided in figure 1. Furthermore, the data can be stored and accessed for longer periods of time without losing any information.[5][6]